Recently, a novel 3D imaging
technique, termed ankylography (derived from the Greek words ankylos meaning curved and graphein meaning writing) has
been developed,
which enables 3D structure determination of a small, general object (or a
relatively large object at lower resolution) from a single view [1]. This work
has ignited a lively debate in the scientific community [2,3]. To
facilitate a better understanding of the method, we post here the Matlab source
codes for the ankylographic reconstructions, and encourage interested readers
to download the codes and test the method. Note that you can use these
source codes to reconstruct any other 3D objects, although the array size is
currently limited. For those who are interested in
the ankylography experiment, two more papers have recently been published [4,5]. In addition, a formal response to the two technical comments
by Wei and Wang et al. has been posted on the arXiv http://arxiv.org/abs/1112.4459. If you
have any questions
about the ankylography method, please
contact John Miao at miao@physics.ucla.edu.
If you have any questions about the source codes, please contact Chien-Chun
Chen at ccchen0627@ucla.edu.
If any of the
following codes are used in your publications and/or presentations, we request
you cite our paper (i.e. ref. 1).
I). Numerical simulation on the ankylographic reconstruction of a 3D "UCLA" pattern with 7 x 7 x 7 voxels (posted on Jan. 20, 2011)
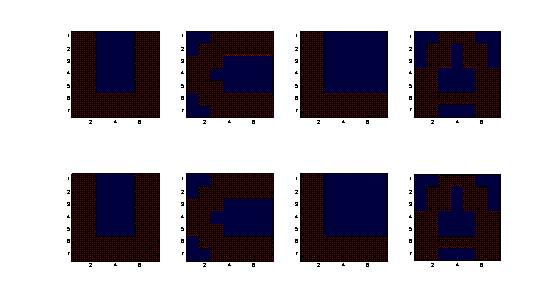
Figure 1. Numerical simulation on ankylographic reconstruction
of a 3D "UCLA" pattern from a spherical diffraction pattern of 1 voxel thick with a
diffraction angle (2q) of 90 degree. The array size of the 3D pattern is 7 x 7 x7 voxels and oversampling degree (Od)
is 1.14 [1]. The upper panel shows the 1st, 3rd, 5th and 7th slices of the reconstructed
image. The lower panel shows the corresponding slices of the 3D pattern,
consisting four alphabet letters "U", "C", "L", and "A".
If you are
interested in reconstructing the 3D "UCLA" pattern, please click
here to download the Matlab
code.
II). Numerical
simulation on the ankylographic reconstruction of a continuous 3D object with
14 x 14 x 14 voxels (posted on Oct. 26, 2011)
![]()
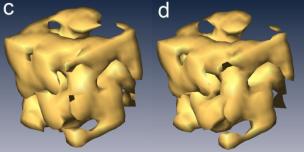
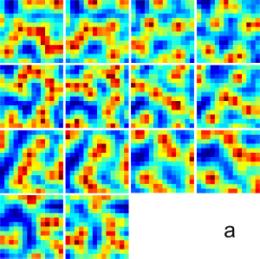
Figure 2. Numerical simulation on the ankylographic reconstruction of a
continuous object with array size of 14 x 14 x 14 voxel. a, 14 slices of the 3D object with a minimum and a maximum voxel
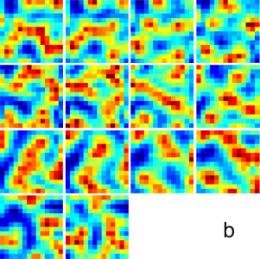
vale of 1.45 and 11.06, respectively. b,
14 reconstructed slices, which are in good agreement with the original ones.
The reconstruction was computed from a simulated spherical different pattern of
1 voxel thick with a diffraction angle (2q) of 90 degree. The oversampling
degree (Od) is 1.48 [1] and
the number of iteration is 106. c,d,
Iso-surface renderings of the original and reconstructed object. The object is
continuous and the holes in the images are due to a threshold value chosen for
the display purpose.
If you are
interested in reconstructing this continuous 3D object, please click
here to download the Matlab
code. In the reconstruction, we usually started with 100 random initial phase
seeds and then chose the best one for the final reconstruction.
III). Numerical simulation on the ankylographic reconstruction of a sodium
silicate glass structure with 25 x 25 x 25 voxels (posted
on Oct. 26, 2011)
![]()
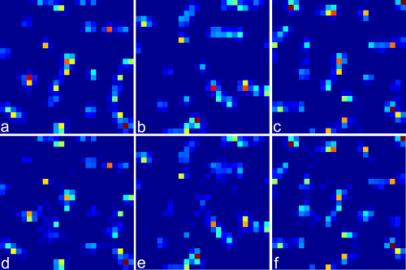
Figure 3. Numerical simulation on the ankylographic reconstruction of a sodium
silicate glass particle. The glass particle structure was generated by molecular
dynamics simulations and consists of a total of 365 atoms with a resolution of
1.5 Angstrom (i.e. 0.75 Angstrom per pixel) and
array size of 25 x 25 x 25 voxels. a-c, Three central slices of the glass
structure along the XY, YZ and XZ planes. d-f,
The corresponding reconstructed slices along the XY, YZ and XZ planes. The
reconstruction was computed from a simulated spherical different pattern of 1
voxel thick with a diffraction angle (2q) of 90 degree. The oversampling degree
(Od) is 1.50 [1] and the
number of iteration is 2x105.
If you are
interested in reconstructing the simulated 3D sodium
silicate glass particle, please click
here to download the Matlab
code. In the reconstruction, we usually started with 100 random initial phase
seeds and then chose the best one for the final reconstruction.
References
1. K.
S. Raines, S. Salha, R. L. Sandberg, H. Jiang, J. A. Rodriguez, B. P. Fahimian,
H. C. Kapteyn, J. Du and J. Miao. Three-dimensional structure determination
from a single view. Natrue 463, 214-217 (2010).
2. P.
Thibault. Feasibility of 3D reconstruction from a single 2D diffraction
measurement, arXiv:0909.1643v1 [physics.data-an] (2009).
3. J.
Miao. Response to "Feasibility of 3D reconstruction from a single 2D
diffraction measurement", arXiv:0909.3500v1 [physics.optics] (2009).
4.C.-C. Chen, H. Jiang, L.
Rong, S. Salha, R. Xu, T. G. Mason and J. Miao. Three-dimensional imaging of a phase object from a single
sample orientation using an optical laser. Phys. Rev. B 84, 224104 (2011).
5. M. D. Seaberg, D. E. Adams,
E. L. Townsend, D. A. Raymondson, W. F. Schlotter, Y. Liu, C. S. Menoni, L. Rong, C.-C. Chen, J. Miao, H. C.
Kapteyn and M. M. Murnane. Ultrahigh 22 nm resolution coherent diffractive imaging using a desktop 13 nm high
harmonic source. Opt. Express 19, 22470-22479 (2011).